Publications
Sabbadin R., Spring D., Rabier C.E.: Dynamic reserve site selection under contagion risk of deforestation (Ecological Modelling, Vol 201, Issue 1, February 2007, p75-81), see also hal-02657333
Likelihood Ratio Test process for Quantitative Trait Locus Detection , with Azais J-M and Delmas C (Statistics, 2012, DOI:10.1080/02331888.2012.760093), see also hal-00483171v4 , authors are in alphabetical order and I am the corresponding author
Data in Statistics : Rice data from Huang et al. (Molecular breeding, 1997)
Rabier C-E, Genz A : The supremum of Chi-Square processes, (Methodology and Computing in Applied Probability, 2013, DOI 10.1007/s11009-013-9331-1), see also hal-00658577
Rabier C-E : On Quantitative Trait Locus mapping with an interference phenomenon, (TEST, Vol 23, 2014, p311-329), see also hal-02632505
Rabier C-E : On statistical inference for selective genotyping, (Journal of Statistical Planning and Inference, Vol 147, April 2014, p24–52), see also hal-00658583v3
Rabier C-E : On empirical processes for quantitative trait locus mapping under the presence of a selective genotyping and an interference phenomenon, (Journal of Statistical Planning and Inference, Vol 153, October 2014, p42-55), see also hal-00796294
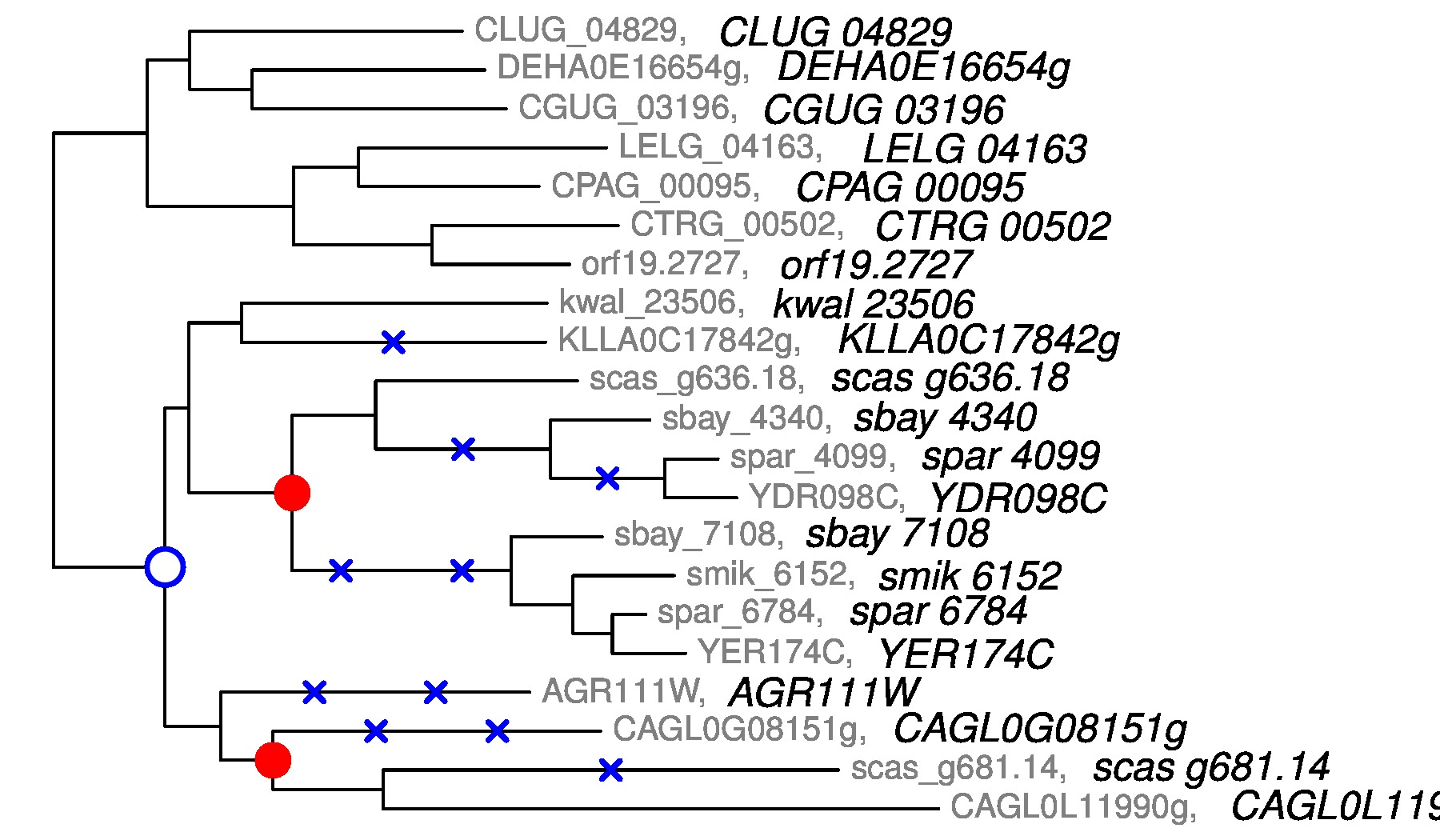
Rabier C-E, Ta T., Ane C. : Detecting and locating Whole Genome Duplications on a phylogeny: a probabilistic approach (Molecular Biology and Evolution, 2014, doi:10.1093/molbev/mst263)
Data in MBE: yeast data, same gene families as in Butler et al. (Nature 2009), that is to say 9209 gene families, downloaded from http://compbio.mit.edu/candida
Rabier C-E : An asymptotic test for Quantitative Trait Locus detection in presence of missing genotypes (Annales de la Faculté des Sciences de Toulouse Mathématiques, Sér 6, Vol 23(4), 2014, p755-778), see also hal-00796296
Rabier C-E : On the asymptotic robustness of the Likelihood Ratio Test in Quantitative Trait Locus detection (Electronic Journal of Statistics, Vol 8(2), 2014, p2138-2157) see also hal-00796295
Rabier C-E : On stochastic processes for Quantitative Trait Locus mapping under selective genotyping (Statistics, Vol 49(1), 2015, p19-34), see also hal-00675414
Rabier C-E, Barre P, Asp T, Charmet G, Mangin B : On the accuracy of genomic selection (PLoS One, Vol 11(6), 2016) , see also hal-01595179
Data in the CROPDL project : raygrass (INRA Lusignan + Denmark, included in Plos One), wheat (SupAgro Montpellier, INRA Clermont Ferrand). Raygrass data available here on dryad.
Rabier C-E, Mangin B, Grusea S : On the accuracy in high dimensional models and its application to genomic selection, (Scandinavian Journal of Statistics, 2018), hal-01456310 Version 2 Version 1
Data in SJS : Rice data from Spindel et al. ( Plos Genetics, 2015 ), available here on dryad
Rabier C-E, Azais J-M, Elsen J-M, Delmas C : Chi-square processes for gene mapping in a population with family structure (Statistical Papers, Vol 60(1), 2019, p 239–271), see also hal-02619068
Data in Statistical Papers : human data from HAPMAP project
Mangin B, Rincent R, Rabier C-E, Moreau L, Goudemand-Dugue E. Training set optimization of genomic prediction with EthAcc outperforms usual accuracy and shows that bigger is not better (Plos One, Vol 14(2), 2019).
Data in this Plos One paper: Sunflower (see Sunrise Project ), Wheat (see BreedWheat project ), Sugar Beet (see AKER project ), Maize (see Amaizing Project )
Rabier C-E, Delmas C : The SgenoLasso and its cousins for selective genotyping and extreme sampling: application to association studies and genomic selection , (Statistics, Vol 55(1), 2021, p18-44) see also hal-02123295
Data in the paper : Rice data from Begum et al. ( Plos One, 2016 ), available here
Rabier C-E, Grusea S : Prediction in high dimensional linear models and application to genomic selection under imperfect linkage disequilibrium , (Journal of the Royal Statistical Society Series C, 2021, doi.org/10.1111/rssc.12496), see also hal-01987222
Data in the paper: Rice data from Spindel et al. ( Plos Genetics, 2015 ), available here on dryad
Rabier C-E, Berry V., Stoltz M, Santos J., Wang W., Glaszmann J.C., Pardi F., Scornavacca C.: On the inference of complex phylogenetic networks by Markov Chain Monte-Carlo, Plos Computational Biology, 2021, available also from hal here
Data : Rice data from Joao Santos, JC Glaszmann (CIRAD) and Wensheng Wang (Chinese Academy of Agricultural Sciences), (raw data from the international rice research institute)
Rabier C-E, Delmas C: The AdaptSgenoLasso, an extended version of the SgenoLasso, for gene mapping and for genomic prediction using the extremes, in press Statistics 2024, hal-04059080
PhD Dissertation (University Toulouse 3, Paul Sabatier, Mathematical Statistics)
Rabier C-E (2010): Techniques statistiques pour la detection de genes a effets quantitatifs
Softwares
SnappNet : SnappNet is a new Bayesian method dedicated to phylogenetic network inference.
It is distributed as a
BEAST package. This project handles the graphical user interface Beauti from BEAST and you do not have to handle xml files. SnappNet's method is described in the submitted paper
``On the inference of complex phylogenetic networks by Markov Chain Monte-Carlo". SnappNet manual can be downloaded
here from github.
Some extra informations are given here . Be careful xml informations are linked to another project
called SnappNetForSimSnappNet (also available on github) that we used for analyzing simulated data.
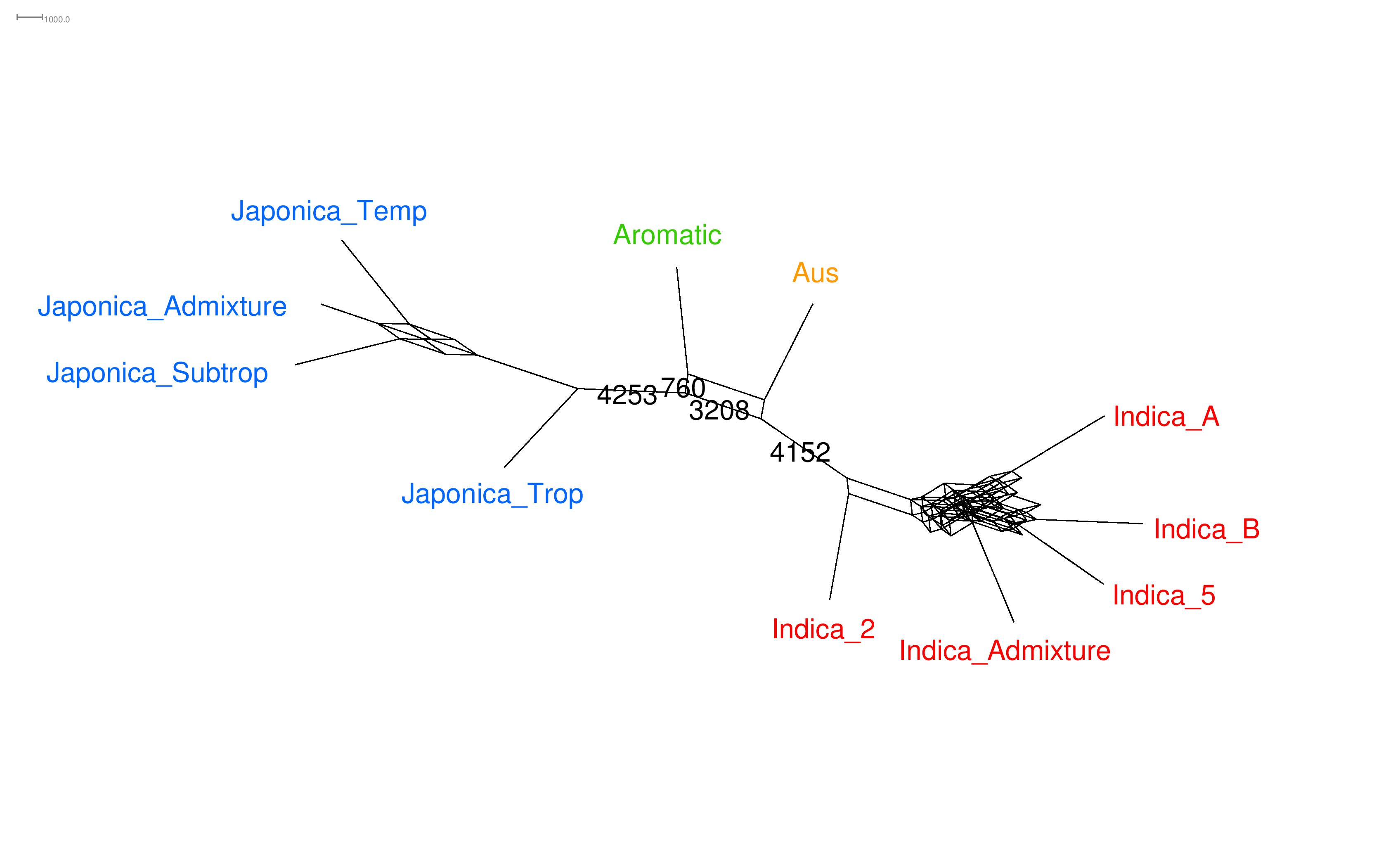
SimSnappNet : SimSnappNet is a c++ simulator that generates count data according to the network multispecies coalescent model.
SimSnappNet is also described here . SimSnappNet is build on the software SimSnapp written by David Bryant and Remco Bouckaert, but it handles now networks.
SgenoLasso : our R code for the paper ``The SgenoLasso and its cousins for selective genotyping and extreme sampling: application to association studies and genomic selection "
GSImperfectLD : our R code for the paper `Prediction in high dimensional linear models and application to genomic selection under imperfect linkage disequilibrium". It requires the
R package hypred of Franck Technow. In order to install hypred,
download it from here , and then you need to recompile the source
in order to make it work with last version of R. So, write in your terminal: R CMD build hypred.
Genomic Selection : our R code for the Scand. J. Stat. paper ``On the accuracy in high dimensional models and its application to genomic selection".
It requires the R package hypred of Franck Technow. See above on how to install hypred.
Genomic Selection : our R code for the Plos One paper ``Training set optimization
of genomic prediction with EthAcc outperforms usual accuracy and shows that bigger is
not better".
Genomic Selection : our R code for the Plos One paper ``On the accuracy of genomic selection". It requires the
R package hypred of Franck Technow. See above on how to install hypred.
SPIMAPWGD and WGDgc : softwares for reconstructing gene trees and detecting Whole Genome Duplications
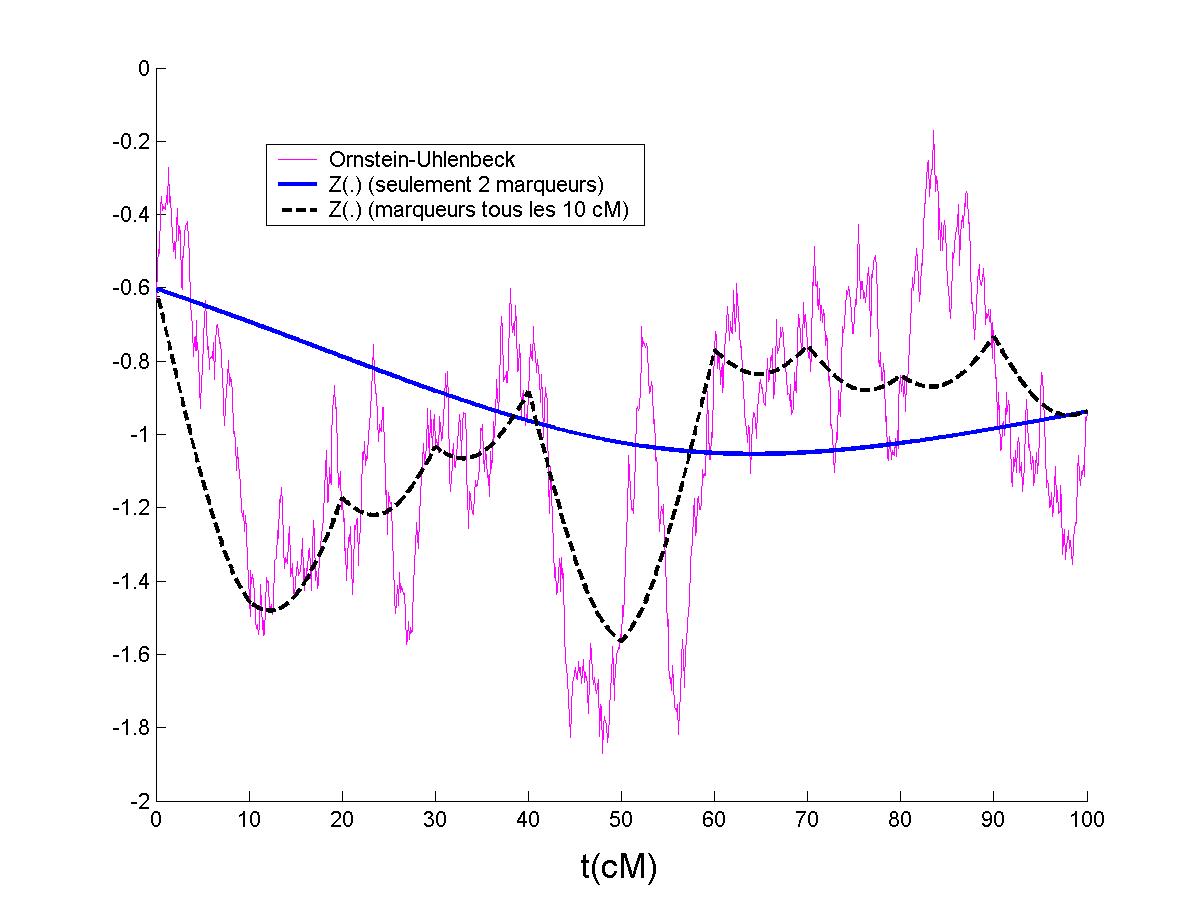
imapping : this is a package with a graphical interface implementing the method presented in the article "Likelihood Ratio Test Process for Quantitative Trait Locus Detection". Download the package and just write pop in your matlab console.
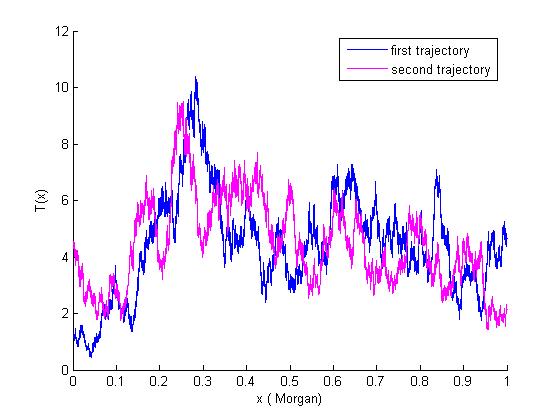
imappingfamily : this is a package with a graphical interface regrouping the different methods presented in the article "Chi-square processes for gene mapping in a population with family structure". Download the package and just write pop in your matlab console.
Invited conferences (underlined speaker)
C Ané, T Ta, C-E Rabier : Probabilistic approaches for detecting and locating whole genome duplications, Statistical Methods for Post-Genomic Data, Paris, France, 2014.
C-E Rabier : Gaussian and Chi-Square processes for Quantitative Trait Locus mapping under selective genotyping, International Indian Statistical Association Conference, Riverside, California, USA, July 2014
B Mangin, C-E Rabier : Training set optimization in genomic selection, 17th meeting of the EUCARPIA Section Biometrics in Plant Breeding, Ghent, Belgium, 2018.
Conferences/Workshops (underlined speaker)
R. Sabbadin, D.Spring, C-E Rabier : Dynamic reserve site selection under contagion risk of deforestation, International Congress on Modelling and Simulation, Melbourne, Australia, 2005.
C-E Rabier, J-M Elsen, C. Delmas : Rejection thresholds in Quantitative Trait Loci detection, 11th QTL-MAS Workshop, Toulouse, 2007
C-E Rabier, J-M Azais : Selective Genotyping for Quantitative Trait Locus detection, Meeting of the MAFIA team of Laboratoire de Statistiques et Probabilites de Toulouse, Nissan lez Enserunes, France, 2007
C-E Rabier, J-M Azais : Selective Genotyping for Quantitative Trait Locus detection, Statistical Meeting, Santander-Toulouse-Valladolid, Valladolid, Spain, 2008
C-E Rabier, J-M Elsen, C. Delmas : On the theory of Quantitative Trait Locus detection, 24th International Biometric Conference, Dublin, Ireland, 2008
C-E Rabier, J-M Elsen, C. Delmas : Rejection thresholds in Quantitative Trait Locus detection, Meeting of the European Association for Animal Production, Vilnius, Poland, 2008
C-E Rabier, J-M Azais : Selective Genotyping for Quantitative Trait Locus detection, Meeting Statistics and its applications, Frejus, France, 2008
C-E Rabier, J-M Azais : Selective Genotyping for Quantitative Trait Locus detection, Meeting of the French Statistical society, Bordeaux, France, 2009
C-E Rabier, J-M Azais, C. Delmas : Likelihood Ratio Test process for the detection of Quantitative Trait Locus, Meeting of the French Statistical society, Marseille, France, 2010
C-E Rabier : My research in QTL detection, Stat/Math Phylogenetics group, University of Wisconsin-Madison, USA, 2011
C-E Rabier, Delmas C : Likelihood Ratio Test process for Quantitative Trait Loci detection, Joint Statistical Meetings, Miami Beach, USA, 2011
C-E Rabier, Ane C : Testing Whole Genome Duplications, Stat/Math Phylogenetics group, University of Wisconsin-Madison, USA, 2012
C-E Rabier, Ta T, Ane C : Testing Whole Genome Duplications, Evolution Seminar Series, University of Wisconsin-Madison, USA, 2013
C-E Rabier, Ta T, Ane C : Testing Whole Genome Duplications, Evolution 2013, Snowbird, USA.
C Ané, Ta T, Rabier C-E : Probabilistic approaches for detecting and locating whole genome duplications, Seminar of the laboratory of Biometry and Evolutive Biology, University Lyon 1, 2014.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Statistics Seminar, Mathematics Research Institute of Rennes, France, 2014.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Statistics and Probability Seminar, University of Nice, France, 2014.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Statistics Seminar, University Joseph Fourier, Grenoble, France, 2014.
C-E Rabier: Gaussian processes and Phylogenetic trees for the genome. Statistics Seminar, Statistics and Genome, University of Evry, France, 2014.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Statistics and Probability Seminar, University of Angers, France, 2014.
C-E Rabier: Gaussian processes and Phylogenetic trees for the genome. Mathematics, Evolution and Genomic Seminar, University Aix-Marseille, France, 2014.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Statistics and Probability Seminar, University of Pau, France, 2014.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Statistics Seminar, University Lyon 1, France, 2014.
C-E Rabier: Ta T, Ane C. Testing Whole Genome Duplications. MIAT Unit Seminar, INRA Toulouse, France, 2014.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Seminar of MODAL team, INRIA Lille, 2015.
C-E Rabier: Ta T, Ane C. Testing Whole Genome Duplications. Seminar of GenPhySE Bionformatics team, INRA Toulouse, France, 2015.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Seminar of Mathematics, University of South Brittany, France, 2015.
C-E Rabier: Gaussian and Chi-Square processes for Quantitative Trait Locus detection. Seminar of Probability and Statistics, University of Bordeaux 1, France, 2015.
C-E Rabier, Ta T, Ane C: Testing Whole Genome Duplications. Seminar of the Methods and Algorithms for Bioinformatics (MAB) group, LIRMM, Montpellier, 2015.
C-E Rabier: Gaussian processes and phylogenetic trees in genomics. Statistics seminar, ENSAI, Rennes, France, 2015.
C-E Rabier , Barre P, Charmet G, Mangin B : On the accuracy of genomic selection. IMS-China International Conference on Statistics and Probability, Kunming, China 2015.
C-E Rabier , Barre P, Charmet G, Mangin B : On the accuracy of genomic selection. MIAT Unit Seminar, INRA Toulouse, France, 2015.
C-E Rabier: Gaussian processes for gene mapping. Statistics seminar, University of Strasbourg, France, 2015.
C-E Rabier, C Delmas: On gene mapping with the mixture model and the extremes. Meeting of the French Statistical society, Montpellier, France, 2016.
C-E Rabier: My research in statistical genetics. Working group Genome Harvest, CIRAD, Montpellier, France, 2016.
C-E Rabier, V. Berry, F. Pardi, C. Scornavacca: Modèles et outils à l'interface entre la génétique des populations et de la phylogénétique pour la phylogénomique du riz. Working group Genome Harvest, CIRAD, Montpellier, France, 2017.
C-E Rabier, V. Berry, F. Pardi, C. Scornavacca: Apport des approches phylogénétiques pour expliquer l'origine des génomes mosaïques, exemple chez le Riz, Genome Harvest day, CIRAD, Montpellier, France, 2017.
C-E Rabier, B.Mangin: On prediction in genomic selection , R2D2 Meeting (working group from INRA), Noirmoutiers, France, 2017.
C-E Rabier: Apprentissage en selection génomique, et reconstruction d'arbres et de réseaux en phylogénie, séminaire de l'équipe Méthodes et Algorithmes pour la Bioinformatique, LIRMM Montpellier, France, 2018.
C-E Rabier, V. Berry, F. Pardi, C. Scornavacca: Apport des approches phylogénétiques pour expliquer l'origine des génomes mosaïques, exemple chez le Riz. 9èmes Journées Francophones sur les Réseaux Bayésiens et les Modèles Graphiques Probabilistes, Toulouse, France, 2018.
C-E Rabier, V. Berry, M. Labous, F. Pardi, C. Scornavacca: Apport des approches phylogénétiques pour expliquer l'origine des génomes mosaïques, exemple chez le Riz. Working group Genome Harvest, CIRAD, Montpellier, France, 2018.
C-E Rabier : Prédiction en grande dimension, extrêmes et réseaux. Séminaire de l'ISAE-Supaero, Toulouse, France, 2018.
C-E Rabier : Approches phylogénétiques pour expliquer l'origine des génomes mosaïques, et prédiction en sélection génomique. Séminaire de l'unité GAFL, INRA d'Avignon, France, 2019.
C-E Rabier , B Mangin, S Grusea : On the accuracy in high dimensional linear models and its application to genomic selection. Highlight (oral presentation), JOBIM, Montpellier, France, 2020. Summary available here.
C-E Rabier, V. Berry, J.C. Glaszmann, F. Pardi, C. Scornavacca: On the inference of complex phylogenetic networks by Markov Chain Monte-Carlo. Poster, JOBIM, Montpellier, France, 2020. Summary available here.
C-E Rabier , V. Berry, J.C. Glaszmann, F. Pardi, C. Scornavacca: Inférence de réseaux phylogénétiques pour la détection d'introgressions et d'hybridations. Séminaire de la KIM Data And Life Sciences, Institut Montpelliérain Alexander Grothendieck, Montpellier, France, 2020.
C-E Rabier , C. Delmas: L'AdaptSgenoLasso, une variante du SgenoLasso, pour la localisation de gènes et la prédiction génomique à l'aide des extrêmes. Journées de la société Francaise de Statistique, Nice, France, 2021. Acte disponible ici
C-E Rabier , C. Delmas: The SgenoLasso and its cousins for selective genotyping and extreme sampling. Highlight (oral presentation), JOBIM, Paris, France, 2021. Summary available here
G. Durif , C. Pierkot, C-E Rabier, S. Tieo : Reproducible research: Good practices and useful information , Montpellier, France, 2021. Available here
G. Durif, C. Pierkot, C-E Rabier , S. Tieo: Reproducible research: R , Montpellier, France, 2021. Available here
G. Durif, C. Pierkot, C-E Rabier, S. Tieo : Reproducible research: Python , Montpellier, France, 2021. Available here
C-E Rabier , V. Berry, J.C. Glaszmann, F. Pardi, C. Scornavacca: Introducing SnappNet, Genome Harvest Final Session, CIRAD, Montpellier, France, 2021.
C-E Rabier , C. Restrepo-Ortiz : Statistical approaches for data mining, Spring School of KIM Food and Health, La Grande Motte, France, 2022.
C-E Rabier : Apprentissage en génomique et reconstruction de réseaux en phylogénie, Séminaire de l'axe probabilités statistiques de l'UMR MISTEA, Montpellier, France, 2022.
C-E Rabier , S. Grusea : Prédiction génomique en grande dimension : densité de marqueurs nécessaire pour une prédiction fiable en sélection génomique, Journées de la société Francaise de Statistique, Lyon, France, 2022. Acte disponible ici , classifié dans la thématique Apprentissage Statistique de la SFDS
C-E Rabier , V. Berry, J.C. Glaszmann, F. Pardi, C. Scornavacca. On the inference of complex phylogenetic networks with SnappNet. Evolution Conference, Cleveland, USA, 2022.
C-E Rabier , C. Delmas : New statistical methods for association studies and genomic prediction. 4th International Conference on Statistics: Theory and Applications , Prague, Czech Republic, 2022. DOI:10.11159/icsta22.135, Proceeding available here, thanks to ASET international , Statistical Methodology session.
C-E Rabier , C. Delmas : The SgenoLasso for gene mapping and genomic prediction, 24th International Conference on Computational Statistics (COMPSTAT 2022), Bologna, Italy, 2022. Machine Learning session.
C-E Rabier , R. Leblois, G. Didier, S. Boitard : Séquences d'arbres, graphes de recombinaison ancestraux, et réseaux phylogénétiques. Workshop du projet DevOCGen (BIODIVOC), Moulis, France, 2023.
C-E Rabier , S. Grusea : Prediction in high dimensional linear models and application to genomic prediction with a sparse genetic map. Statlearn'23, Poster Session, Montpellier, France, 2023, available here.
C-E Rabier , C. Delmas : The SgenoLasso, a new Lasso method dedicated to extreme observations in genomics , International Conference on Robust Statistics (ICORS 2023), Toulouse, France, 2023.
C-E Rabier , R. Leblois, G. Didier, S. Boitard : Tree sequences, recombination ancestral graphs, and phylogenetic networks. Evolution Conference, USA, 2023, cancelled.
C-E Rabier : Genomic prediction and Phylogenetics. University of Angers, Seminar at IRHS, 2024.
C-E Rabier , V Berry, J-C Glaszmann : Phylogenetic network inference with ABC Random Forest : application to rice domestication. Mathematical and Computational Evolutionary Biology 2025, Granada, Spain, poster session, 2025.
C-E Rabier , V Berry, J-C Glaszmann : Inférence de réseaux phylogénétiques par ABC-Forêts Aléatoires (ABC-RF) : application au processus de domestication du riz, Journées de la Société Française de Statistique, Marseille, 2025.
A Metzger, K Raschel, C-E Rabier : Pangénomique et réseaux phylogénétiques (Projet Pan-Phylo-Net). Université d'Angers, 2025.
C-E Rabier , C Delmas : The AdaptSgenoLasso for gene mapping and genomic prediction. 2025 IMS International Conference on Statistics and Data Science (ICSDS), Sevilla, Spain, 2025
Webdesign service by Sarkis. Outsourcing by FreelanceWebmarket.